As the world is seeing the plague of COVID-19, an illness brought about by a novel coronavirus, SARS-CoV-2, developing hereditary qualities and clinical confirmations recommend a comparative way to those of SARS and MERS. A course of viral particles enters the body through the nose, eyes or mouth. Breathing conveys a portion of these particles to the lower respiratory tract where the spike proteins of the coronavirus, acting like a key, lock into epithelial cells that line the respiratory tract just as those in the air sacs in the lungs. SARS-CoV-2 can remain undetected longer than numerous influenza or coronaviruses and its spike proteins can pick up passage by opening the ACE2 protein on the lung cells. People might be tainted by and experience the ill effects of various distinctive infections, and in many occasions the disease is settled with or without tissue harm. Reinfection is normally subclinical, and for some infections we have compelling immunizations; exemplary models incorporate measles, mumps, rubella, rotavirus and varicella zoster infections. Different infections, for example, HIV, hepatitis C infection (HCV), hepatitis B infection (HBV) and some herpesviruses, can cause significant tissue harm in a few or all people they contaminate, and sores can get incessant. These infections as a rule have at least one properties that permit them to decrease the viability of host versatile or natural resistance, and we need powerful immunizations against the vast majority of these specialists. Contamination with infections, for example, flu infection and respiratory syncytial infection (RSV) has a variable result. Most people may endure mellow or subclinical contamination, yet others experience serious ailment that can be deadly. Numerous infections taint people and most are controlled agreeably by the resistant framework with restricted harm to have tissues. Some infections, in any case, harm the host, either in segregated cases or as a response that ordinarily happens after disease. The result is affected by properties of the tainting infection, the conditions of contamination and a few variables constrained by the host. A coronavirus is a sort of basic infection that causes a disease in your nose, sinuses, or upper throat. Most coronaviruses aren't perilous. COVID-19 is an illness that can cause what specialists call a respiratory tract disease. It can influence your upper respiratory tract (sinuses, nose, and throat) or lower respiratory tract (windpipe and lungs). It's brought about by a coronavirus named SARS-CoV-2.
Corona virus, COVID-19, Immune response, Infection
Coronaviruses are a group of related RNA viruses that cause diseases in mammals and birds. In humans, these viruses cause respiratory tract infections that can range from mild to lethal. Mild illnesses include some cases of the common cold (which is caused also by certain other viruses, predominantly rhinoviruses), while more lethal varieties can cause SARS, MERS, and COVID-19. Symptoms in other species vary: In chickens, they cause an upper respiratory tract disease, while in cows and pigs they cause diarrhea. There are as yet no vaccines or antiviral drugs to prevent or treat human coronavirus infections [1-4].
Coronaviruses constitute the subfamily Orthocoronavirinae, in the family Coronaviridae, order Nidovirales, and realm Riboviria [5,6]. They are enveloped viruses with a positive-sense single-stranded RNA genome and a nucleocapsid of helical symmetry. The genome size of coronaviruses ranges from approximately 26 to 32 kilobases, one of the largest among RNA viruses [7]. They have characteristic club-shaped spikes that project from their surface, which in electron micrographs create an image reminiscent of the solar corona, from which their name derives [8] (Figure 1 and Figure 2).
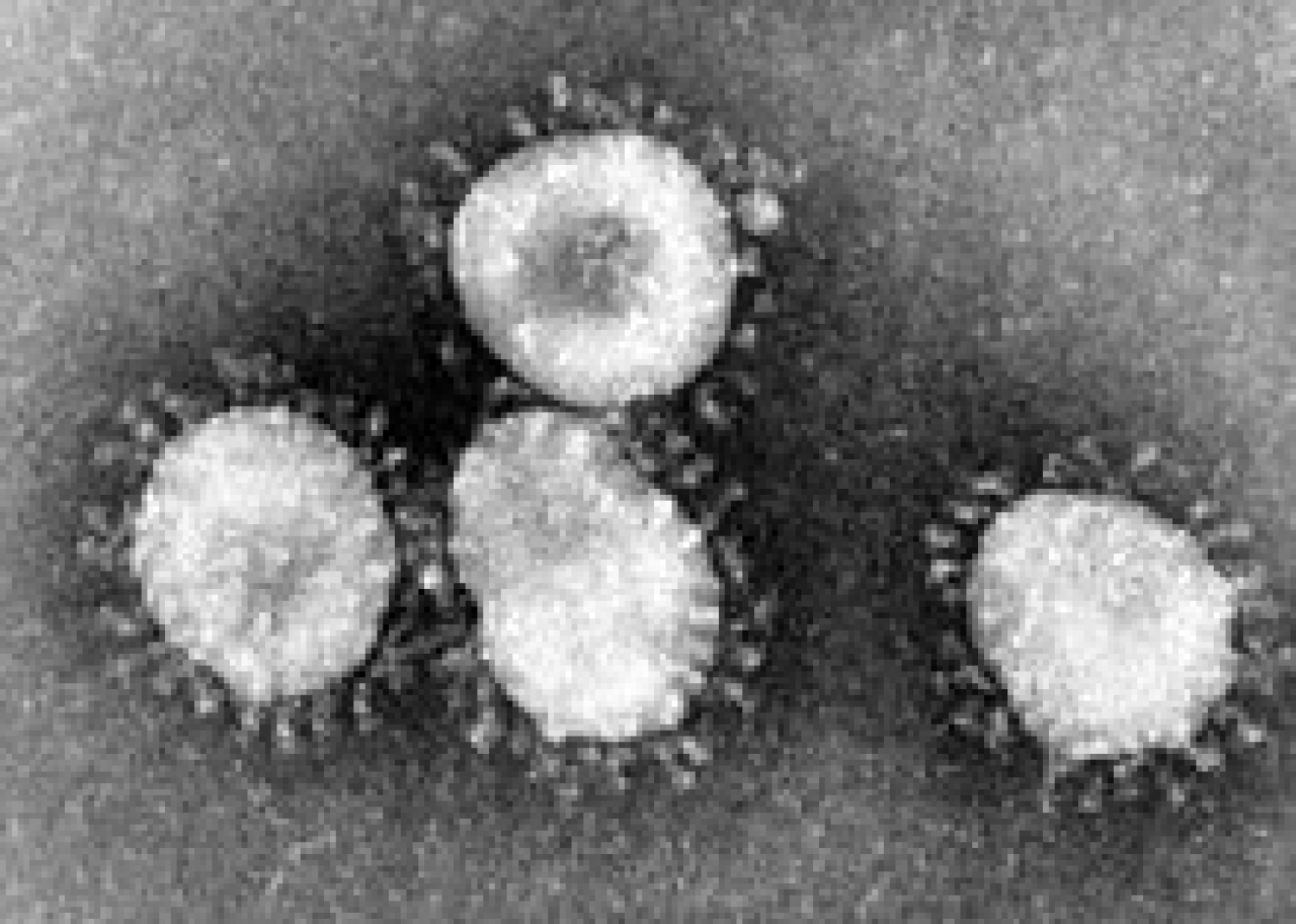 Figure 1: Transmission electron micrograph (TEM) of avian infectious bronchitis virus.
View Figure 1
Figure 1: Transmission electron micrograph (TEM) of avian infectious bronchitis virus.
View Figure 1
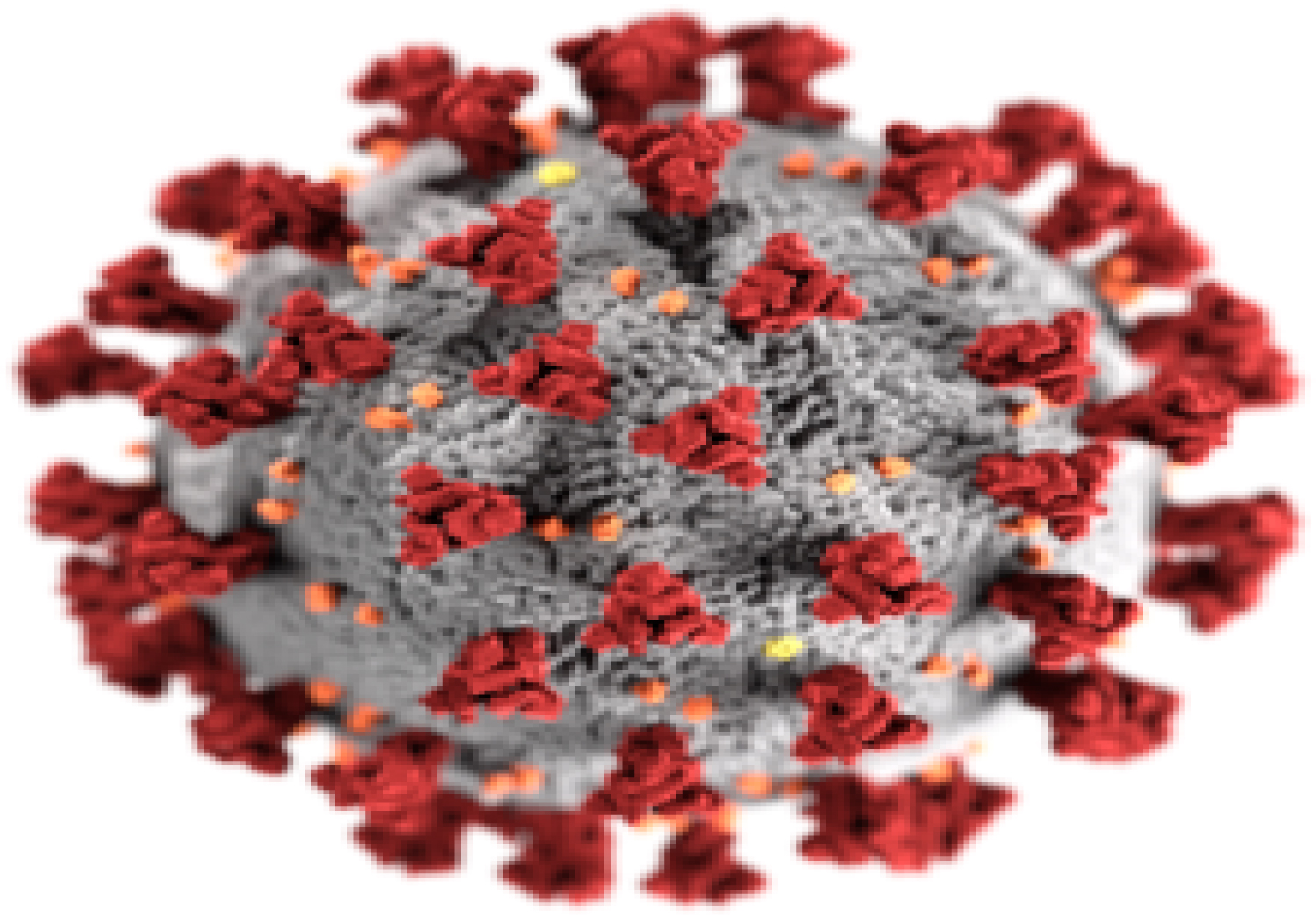 Figure 2: Illustration of the morphology of coronaviruses; the club-shaped viral spike peplomers, colored red, create the look of a corona surrounding the virion when observed with an electron microscope.
View Figure 2
Figure 2: Illustration of the morphology of coronaviruses; the club-shaped viral spike peplomers, colored red, create the look of a corona surrounding the virion when observed with an electron microscope.
View Figure 2
Coronaviruses were first discovered in the 1930s when an acute respiratory infection of domesticated chickens was shown to be caused by infectious bronchitis virus (IBV) [9]. Arthur Schalk and M.C. Hawn described in 1931 a new respiratory infection of chickens in North Dakota. The infection of new-born chicks was characterized by gasping and listlessness. The mortality rate of the chicks was 40-90% [10]. Fred Beaudette and Charles Hudson six years later successfully isolated and cultivated the infectious bronchitis virus which caused the disease [11]. In the 1940s, two more animal coronaviruses, mouse hepatitis virus (MHV) and transmissible gastroenteritis virus (TGEV), were isolated [12]. It was not realized at the time that these three different viruses were related [13].
Human coronaviruses were discovered in the 1960s [14,15]. They were isolated using two different methods in the United Kingdom and the United States [16]. E.C. Kendall, Malcom Byone, and David Tyrrell working at the Common Cold Unit of the British Medical Research Council in 1960 isolated from a boy a novel common cold virus B814 [17-19]. The virus was not able to be cultivated using standard techniques which had successfully cultivated rhinoviruses, adenoviruses and other known common cold viruses. In 1965, Tyrrell and Byone successfully cultivated the novel virus by serially passing it through organ culture of human embryonic trachea [20]. The new cultivating method was introduced to the lab by Bertil Hoorn [21]. The isolated virus when intranasally inoculated into volunteers caused a cold and was inactivated by ether which indicated it had a lipid envelope [17,22]. Around the same time, Dorothy Hamre and John Procknow at the University of Chicago isolated a novel cold virus 229E from medical students, which they grew in kidney tissue culture. The novel virus 229E, like the virus strain B814, when inoculated into volunteers caused a cold and was inactivated by ether [23].
By now, researchers and health experts have gained a better understanding of the range of symptoms caused by COVID-19, which include fever, a dry cough, and of course, the dangerous inflammation of the respiratory system. Most of us know that COVID-19 can be much more severe than a typical flu, but lesser known are the mechanics behind how the virus causes pneumonia in its victims. According to the World Health Organization (WHO), most people who contract COVID-19 only experience mild flu-like symptoms. Occasionally though, the infection can cascade into a severe case of pneumonia that can be lethal, especially for older people and those with underlying medical conditions (Figure 3) [24].
 Figure 3: Occasionally, the infection can cascade into a severe case of pneumonia that can be lethal [24].
View Figure 3
Figure 3: Occasionally, the infection can cascade into a severe case of pneumonia that can be lethal [24].
View Figure 3
The virus, officially named SARS-CoV-2, enters the body - generally through the mouth or nose. From there, the virus makes its way down into the air sacs inside your lungs, known as alveoli.
Once in the alveoli, the virus uses its distinctive spike proteins to "hijack" cells. The primary genetic programming of any virus is to make copies of itself, and COVID-19 is no exception. Once the virus' RNA has entered a cell, new copies are made and the cell is killed in the process, releasing new viruses to infect neighboring cells in the alveolus.
The process of hijacking cells to reproduce causes inflammation in the lungs, which triggers an immune response. As this process unfolds, fluid begins to accumulate in the alveoli, causing a dry cough and making breathing difficult. For 80-85% of people infected by COVID-19, these symptoms will run their course much as they would with a case of the flu.
In 15-20% cases, the immune system's response to inflammation in the lungs can cause what's known as a "cytokine storm". This runaway response can cause more damage to the body's own cells than to the virus it's trying to defeat, and is thought to be the main reason for why the conditions of young, otherwise healthy individuals can rapidly deteriorate.
If enough alveoli collapse, acute respiratory distress syndrome (ARDS) can occur, requiring a patient to be placed on a ventilator for breathing assistance (Figure 4).
 Figure 4: In 15-20% cases the immune system's response can cause a 'cytokine storm' [24].
View Figure 4
Figure 4: In 15-20% cases the immune system's response can cause a 'cytokine storm' [24].
View Figure 4
At this stage, the surfactant that helps keep alveoli from collapsing has been diluted, and fluid containing cellular debris is impairing the gas exchange process that supplies oxygen to our bloodstream.
In the most severe cases, systemic inflammatory response syndrome (SIRS) occurs as the protein-rich fluid from the lungs enters the bloodstream, resulting in septic shock and multi-organ failure. This is often the cause of death for people who have succumbed to a COVID-19 infection.
However, in reality, the physiological processes underlying these phases overlap. People with COVID-19 may or may not show features of earlier or later phases [25].
Both SARS-CoV-2 and SARS-CoV gain entry via a receptor called ACE2. More commonly known for their role in controlling blood pressure and electrolytes, these receptors are also present in the lungs, back of the throat, gut, heart muscle, and kidneys. In 2004, researchers from the University Medical Center Groningen in the Netherlands reported that ACE2 receptor cells were not present on the surface layer of cells in the nose and were, therefore, not an important site for SARS-CoV viral replication. In SARS, there are hardly any upper respiratory tract symptoms, and viral units are rarely present outside the lungs. This fact initially took the focus away from continuing to look for ACE2 receptors in the nose.
Recently, an international team of researchers has found the ACE2 receptors on goblet (secretory) cells in and on ciliated (hairy) cells in the nose. More recently, scientists have found ACE2 receptors in the mouth and tongue, potentially indicating a hand-to-mouth route of transmission. Researchers also found a plentiful supply of a protease called TMPRSS2, which chemically splits off the top of the coronavirus spike to allow the SARS-CoV-2 RNA to enter into the nasal cells. Once inside the cell, the virus's genetic material directs the cell to manufacture millions of new copies of itself.
As a result, SARS-CoV-2 can bind 10 times more tightly to insert its RNA into the cell, starting to explain why COVID-19 spreads so rapidly [25,26] (Figure 5).
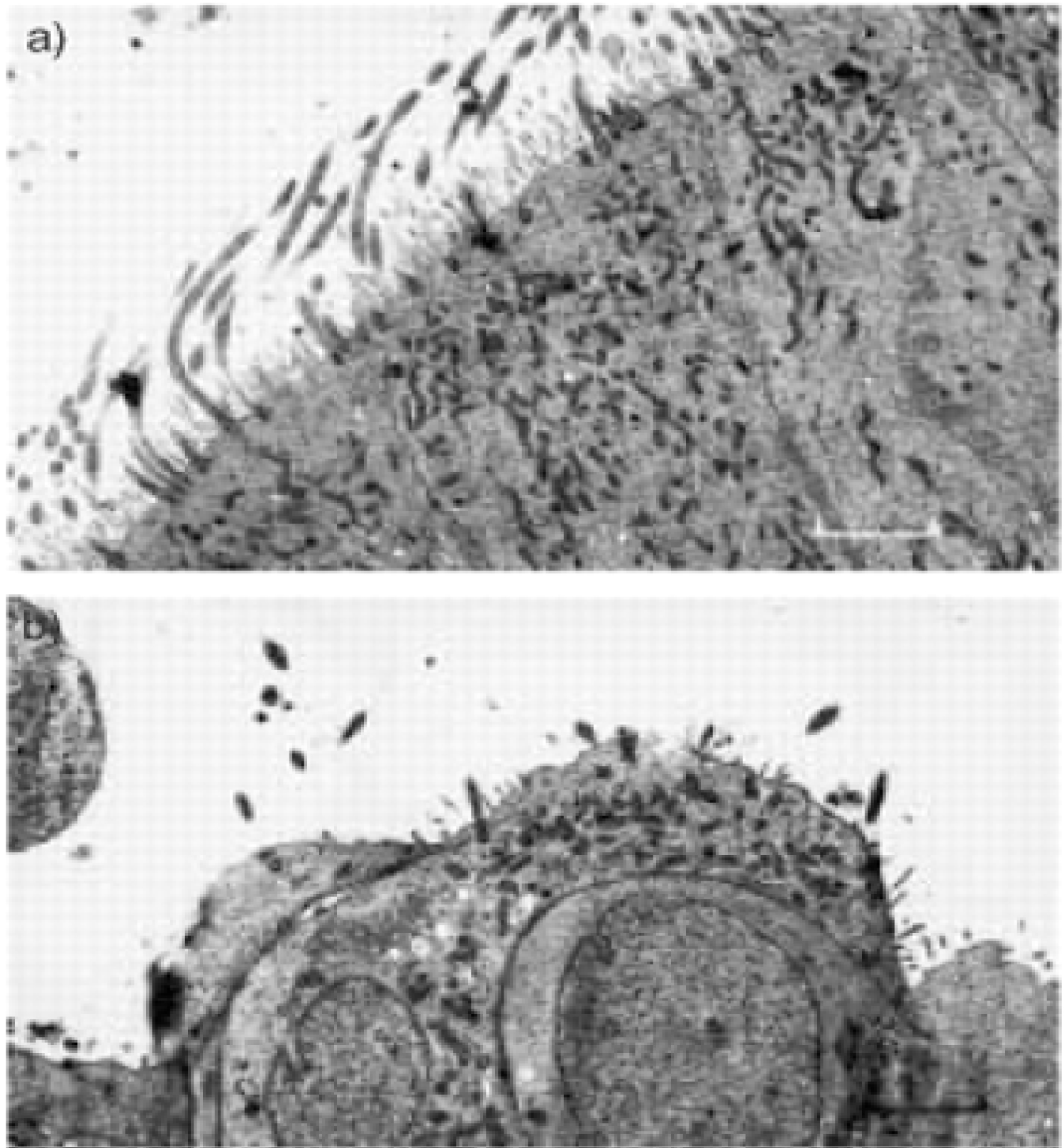 Figure 5: Transmission electron micrograph of nasal epithelium before and after coronavirus inoculation. a) Transmission electron micrograph day 0. This shows normal tissue with an intact well-ciliated surface and minimal disruption; b) Transmission electron micrograph day 3. This shows abnormal tissue with severely disrupted cell surface. Marked loss of cilia is seen. Internal scale bars = 2.9 µm [26].
View Figure 5
Figure 5: Transmission electron micrograph of nasal epithelium before and after coronavirus inoculation. a) Transmission electron micrograph day 0. This shows normal tissue with an intact well-ciliated surface and minimal disruption; b) Transmission electron micrograph day 3. This shows abnormal tissue with severely disrupted cell surface. Marked loss of cilia is seen. Internal scale bars = 2.9 µm [26].
View Figure 5
The viral load study in Germany showed that active viral replication occurs in the upper respiratory tract. Seven out of nine participants listed a cough among their initial symptoms. In contrast to the falling numbers of viral units in the upper respiratory tract, numbers in sputum rose for most of the participants. In two individuals with some signs of lung infection, the virus in sputum peaked at day 10-11. It was present in the sputum up to day 28 in one person. Across all participants, there was an average of 7 million units in 1 milliliter (about 35 million units in a teaspoon). This amount is about 1,000 times more than that in people with SARS. In the lung, the ACE2 receptor sits on top of lung cells called pneumocytes. These have an important role in producing surfactant - a compound that coats the air sacs (alveoli), thus helping maintain enough surface tension to keep the sacs open for the exchange of oxygen and carbon dioxide. As soon as the body recognizes a foreign protein, it mounts the first response. One part of the body's immune response - the lymphocytes - begin to produce the first defense IgM-type antibodies and then the longer term specific neutralizing antibodies (the IgG type) [25].
Approximately 13.8% of people with COVID-19 will have severe disease and will require hospitalization as they become short of breath. Of these individuals, 75% will have evidence of bilateral pneumonia. Pneumonia in COVID-19 occurs when parts of the lung consolidate and collapse. Reduced surfactant in the alveoli from the viral destruction of pneumocytes makes it difficult for the lungs to keep the alveoli open.
As part of the immune response, white blood cells, such as neutrophils and macrophages, rush into the alveoli. Meanwhile, blood vessels around the air sacs become leaky in response to inflammatory chemicals that the white blood cells release. This fluid puts pressure on the alveoli from outside and, in combination with the lack of surfactant, causes them to collapse. As a result, breathing becomes difficult, and the surface area in the lung where oxygen transfer usually takes place becomes reduced, leading to breathlessness [25].
The most common time for the onset of critical disease is 10 days, and it can come on suddenly in a small proportion of people with mild or moderate disease. In severe acute respiratory distress syndrome (ARDS), the inflammation stage gives way to the fibrosis stage. Fibrin clots form in the alveoli and fibrin-platelet microthrombi (small blood clots) pepper the small blood vessels in the lung that are responsible for gas exchange with the alveoli. There is hope that drugs already licensed for anticlotting action in strokes could be helpful at this stage. Cytokines are chemical mediators that white blood cells such as macrophages release, and they can engulf infected cells. These cytokines - which have names such as IL1, IL6, and TNFα - have actions that include dilating the vessel walls and making them more permeable. In extreme circumstances, this can lead to a collapse of the cardiovascular system [25].
MERS-CoV and SARS-CoV are β-coronaviruses that can cause deadly lower respiratory tract contaminations and extrapulmonary manifestations [27-29]. T cells, CD4+ T cells, and CD8+ T cells especially assume a critical antiviral job by adjusting the battle against pathogens and the danger of creating autoimmunity or overpowering inflammation [30]. CD4+ T cells advance the creation of virus-specific antibodies by actuating T-dependent B cells. Be that as it may, CD8+ T cells are cytotoxic and can murder viral tainted cells. CD8+ T cells represent about 80% of all-out infiltrative fiery cells in the pneumonic interstitium in SARS-CoV-infected patients and assume an essential job in clearing CoVs in tainted cells and prompting safe injury [31]. What's more, by contrasting T-cell-deficient BALB/c mice (transduced by ad5-hdp4) with controls and B‐cell‐deficient mice, a few specialists verified that T cells could get by in the contaminated lungs and devastate the tainted cells [32]. It underlines the significant job of T cells as opposed to B cells in the control of the pathogenesis of MERS-CoV disease. A cross-reactive T cell reaction prompts a diminishing in MERS‐CoV [33]. However, CD4+ T cells are increasingly defenseless to MERS-CoV contamination. The consumption of CD8+ T cells doesn't influence and postpone viral replication at the hour of contamination with SARS-CoV [34,35]. Depletion of CD4+ T cells is related to a diminished aspiratory enrollment of lymphocytes and killing immunizer and cytokine creation, bringing about a solid immune‐mediated interstitial pneumonitis and deferred freedom of SARS-CoV from lungs [36]. Additionally, T aide cells produce proinflammatory cytokines through the NF-kB flagging pathway [37]. IL-17 cytokines select monocytes and neutrophils to the site of disease with irritation and actuate other downstream cytokine and chemokine falls, for example, IL-1, LL-6, IL-8, IL-21, TNF-β, and MCP-1 [38,39].
B cell subsets with phenotypes normal for credulous, non-isotype-switched, memory cells, and antibody-secreting cells collect in CoVs [40]. The antigen incitement of MERS-CoV disease was explained by utilizing the particular 9-mer peptide "CYSSLILDY", which situated at position 437 to 445 inside the area of the S glycoprotein [40]. The arrangement has the most noteworthy B cell antigenicity plot and can frame the best number of connections with MHCI alleles in an electronic simulation [41]. Reports show that humoral invulnerability is basic to control the diligent period of CoV contamination. More antibodies confined from patients who have to endure MERS-CoV contamination have been portrayed, including MCA1, CDC-C2, CSC-C5, CDC-A2, CDC-A10, MERS-GD27, and MERS-GD33 [42,43].
For most people, Covid-19 seems to be pretty mild. And it takes a while - to the tune of five days to two weeks - to cause symptoms, if it does at all.
Unfortunately, that's exactly what makes the novel coronavirus so dangerous. In the period that an infected person is asymptomatic or mildly ill, they could transmit the virus to dozens of other people through water droplets expelled by coughs or sneezes, transferred on skin and other surfaces. Angiotensin converting enzyme 2 receptors, or ACE2 receptors for short, is a part of the sympathetic nervous system. Their job is to bind to a hormone (conveniently called the ACE2 hormone) as part of the body's stress response. The ACE2 hormone plays a role in constricting blood vessels, which raises blood pressure.
The virus that causes Covid-19 can latch onto the ACE2 receptor undetected - likely because of its animal origins. The adaptive immune system - the part that creates antibodies that help identify pathogens quickly - hasn't had a chance to learn what SARS-CoV-2 looks like yet. This means the virus can use a molecular phishing scam to sneak into the cell. People with diabetes or high blood pressure - who are typically older adults - often take a class of drugs called ACE inhibitors. These drugs limit the constriction of blood vessels and, in doing so, make ACE2 receptors more prevalent throughout the body. Some scientists hypothesize that this is why people with these conditions have much more severe cases of Covid-19.
Although infection with the emerging SARS-CoV was associated with a severe acute respiratory disease, most human CoVs are responsible for mild upper respiratory tract infections, such as common colds, with only occasional spreading to the lower respiratory tract. Most respiratory viruses interact with DCs in the upper respiratory tract, which results in initiating an antiviral immune response but may also result in the spreading of the virus as a result of DC migration to the draining lymph nodes.